Which enzyme unwinds the DNA double helix during replication?
DNA polymerase
Ligase
Helicase
Primase

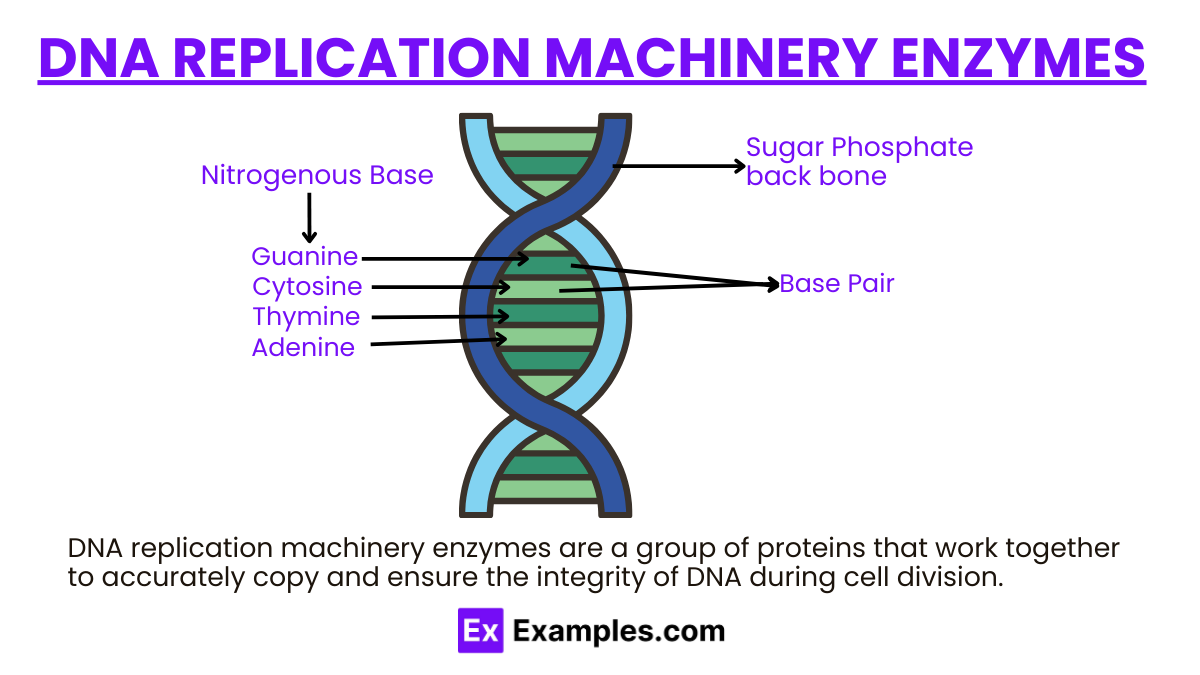
The heart of cellular biology with our comprehensive guide on DNA Replication Machinery Enzymes. This intricate process is the cornerstone of genetic inheritance, involving a suite of enzymes that orchestrate the replication of DNA, ensuring life’s continuity. From the unwinding action of helicase to the synthesizing prowess of DNA polymerase, each enzyme plays a pivotal role. Through detailed examples, this guide sheds light on these molecular maestros, offering a window into the mechanisms that drive biological replication and heredity. Enhance your understanding of the fundamental processes that underpin all living organisms, enriched with keyword-rich content that’s both SEO and NLP friendly, making it a perfect resource for students, professionals, and curious minds alike.
DNA replication is a fundamental process by which a cell duplicates its DNA, ensuring that each new cell receives an exact copy of the original genetic material. This process is critical for cell division, allowing organisms to grow, repair damaged tissues, and reproduce.
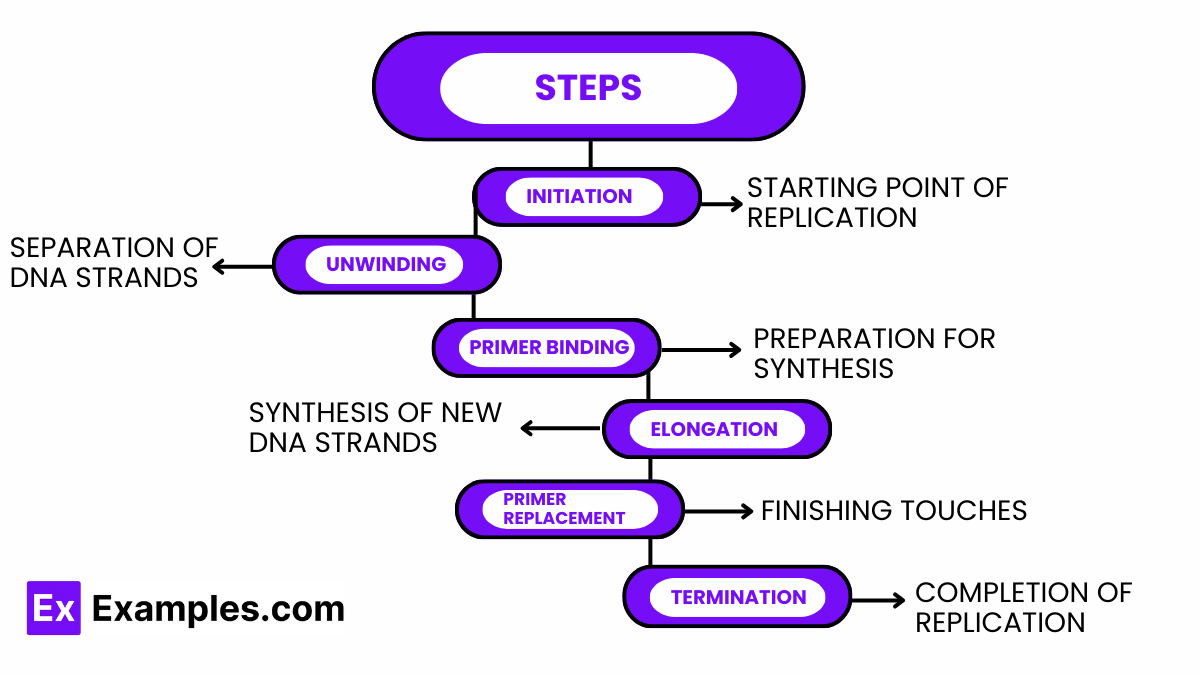
DNA replication is a critical biological process that ensures the transmission of genetic information from one cell generation to the next. It involves a series of orchestrated steps that result in the duplication of the DNA molecule. Below, the key steps of DNA replication are explained in detail.
Starting Point of Replication
Separation of DNA Strands
Preparation for Synthesis
Synthesis of New DNA Strands
Finishing Touches
Completion of Replication
DNA replication is a complex process requiring the orchestrated action of several enzymes, each performing a specific function to ensure accurate and efficient duplication of the DNA molecule. Here’s how these enzymes contribute to DNA replication:
Unwinding the DNA Helix
Synthesis of New DNA Strands
Primer Synthesis
Joining DNA Fragments
Preventing Over-winding
Stabilizing Single Strands
RNA Primer Removal
Primer Replacement
DNA replication in prokaryotes is the process by which a prokaryotic cell duplicates its single, circular chromosome, ensuring that each daughter cell receives a complete copy of the genetic material. This essential biological process involves several key steps:
DNA replication in eukaryotes is a complex and highly regulated process that ensures the accurate duplication of the organism’s much larger and more complex genome, which is organized into multiple linear chromosomes. This process is essential for cell division, allowing each daughter cell to receive a complete set of genetic information. Here are the key points outlining how DNA replication occurs in eukaryotic cells:
| Aspect | DNA Replication | Transcription |
|---|---|---|
| Purpose | To duplicate the entire DNA molecule for cell division. | To produce an RNA copy of a gene for protein synthesis. |
| Location | Occurs in the nucleus (in eukaryotes). | Occurs in the nucleus (in eukaryotes) for mRNA synthesis. |
| Enzymes Involved | Main enzyme is DNA polymerase. | Main enzyme is RNA polymerase. |
| Template | Uses both strands of DNA as templates. | Uses only one strand of DNA as a template. |
| Product | Produces two identical DNA molecules. | Produces a single strand of RNA. |
| Primers Required | Requires primers to initiate replication. | Does not require primers for initiation. |
| Direction | Bidirectional from the point of origin. | Unidirectional, moving from the promoter to the terminator. |
| Processivity | Highly processive, copying long sequences without stopping. | Less processive, synthesizes shorter RNA molecules. |
| Error Rate | Lower error rate due to proofreading mechanisms. | Higher error rate, with limited proofreading. |
| Strand Complementarity | Both new DNA strands are complementary to their template strands. | The RNA strand is complementary to the DNA template strand, but uracil replaces thymine |
DNA replication machinery enzymes are specialized proteins that facilitate the process of DNA replication, where the DNA molecule is duplicated before cell division. These enzymes work together to unwind the DNA double helix, synthesize new DNA strands, and ensure the accuracy of the replication process.
The enzyme that initiates DNA replication is DNA helicase. It unwinds the double-stranded DNA by breaking the hydrogen bonds between the nucleotide bases, creating a replication fork where other replication enzymes can act.
DNA polymerase plays a critical role in DNA replication by adding nucleotides to the growing DNA strand. It reads the template strand and incorporates complementary nucleotides, ensuring the new strand is an accurate copy of the template strand. It also has proofreading capabilities to correct errors.
RNA primers are removed by DNA polymerase I in prokaryotes and by RNase H and FEN1 (Flap Endonuclease 1) in eukaryotes. After the removal, the gaps left by the primers are filled with DNA nucleotides.
DNA ligase seals nicks in the DNA backbone, joining Okazaki fragments on the lagging strand and ensuring the continuity of the newly synthesized DNA strands. This action is crucial for completing the replication process.
Primase synthesizes short RNA primers that are necessary for DNA polymerases to begin DNA synthesis. Since DNA polymerases can only add nucleotides to an existing strand, primase provides the starting point for DNA synthesis.
No, DNA polymerase cannot start DNA synthesis on its own because it requires a primer with a free 3′ hydroxyl group to which it can add nucleotides. This is why primase is essential for initiating DNA replication.
The accuracy of DNA replication is ensured by the proofreading function of DNA polymerase. If an incorrect nucleotide is incorporated, the enzyme can remove it and replace it with the correct one. This reduces the error rate significantly.
The replication fork is formed when DNA helicase unwinds the DNA double helix, separating the two strands. This creates a Y-shaped structure where DNA replication can occur, with one strand serving as the template for leading strand synthesis and the other for lagging strand synthesis.
During DNA replication, telomeres, the repetitive DNA sequences at the ends of eukaryotic chromosomes, tend to shorten. This is mitigated by the enzyme telomerase, which extends the telomeres, preventing loss of important genetic information and maintaining chromosome stability.
In sum, DNA replication machinery enzymes are vital for the accurate duplication of genetic material, ensuring the continuity of life across generations. Through the coordinated actions of helicases, polymerases, primases, and ligases, DNA is replicated efficiently and precisely. Understanding these enzymes sheds light on the fundamental processes that underpin cellular biology and genetic inheritance.
Text prompt
Add Tone
Role of Enzymes in DNA Replication
Difference between Replication and Transcription
Which enzyme unwinds the DNA double helix during replication?
DNA polymerase
Ligase
Helicase
Primase
What is the primary role of DNA polymerase?
Unwinding the DNA
Synthesizing RNA primers
Adding nucleotides to the growing DNA strand
Sealing nicks in the DNA
Which enzyme is responsible for synthesizing RNA primers during DNA replication?
Helicase
Ligase
Primase
Topoisomerase
The enzyme that relieves the tension created by the unwinding of the DNA double helix is:
Helicase
Ligase
DNA polymerase
Topoisomerase
Which of the following enzymes seals the nicks between Okazaki fragments on the lagging strand?
Helicase
DNA polymerase
Primase
Ligase
What is the function of single-strand binding proteins (SSBs) in DNA replication?
Unwinding the DNA
Synthesizing new DNA
Stabilizing the single-stranded DNA
Removing RNA primers
During DNA replication, which enzyme removes RNA primers and replaces them with DNA?
Helicase
Primase
Ligase
DNA polymerase I
Which enzyme initiates the synthesis of a new DNA strand by creating a short RNA primer?
DNA polymerase
Helicase
Ligase
Primase
The leading strand is synthesized:
Discontinuously
In short fragments
Continuously
By RNA polymerase
Okazaki fragments are found on the:
Leading strand
Lagging strand
RNA strand
Single-stranded DNA
Before you leave, take our quick quiz to enhance your learning!

